|
|

Fossils, Earth History May Hold Clues For New Cures
The human genome contains 3 billion letters of DNA sequence and as many as 100,000 genes. Buried within this enormous pile of data is information about why people get sick and what might be done to keep them healthy. Finding this information, however, has proved difficult, with the result that the human genome project has not yet proved to be the medical windfall that was long anticipated. In an article in the journal Science in May, Steven Benner, a distinguished professor of chemistry, wrote that a solution may lie in connecting the history of genes and proteins to the geological record and the fossil records of our human and pre-human past. "The past is key to the present," Benner says. "By understanding how proteins evolved together with the history of the planet, we can better understand how proteins work today." Proteins control life functions. To find how they evolved, molecular geneticists extract information about their ancestors by matching up similarities in gene sequences. Geneticists seek similarities in amino acids denoted by the different letters of the genetic code. They use these similarities to reconstruct the amino acid sequences of ancient proteins and resurrect these ancient proteins in the laboratory, studying them to gain insights into their modern descendants. The next step is to date the events that produced the proteins and the genes associated with them. The Science paper reports a new way to date events recorded in genomes from sequence data alone. Benner conducted the research in collaboration with scientists at UF and the Foundation for Applied Molecular Evolution in Gainesville. With the dates in hand, scientists can then match the events that created genes and proteins with events recorded in the fossil and geological records. This allows scientists to zero in on the genes and proteins they may be hunting, Benner says. As an example of how this approach may benefit the search for therapies, Benner cites the cautionary tale of leptin, the so-called "obesity gene." Rockefeller University scientists discovered this gene in mice in 1994. When they deleted it, mice became obese, creating widespread excitement that the parallel gene in humans would perform the same function, opening the door to genetic therapy or cure for obesity. That goal has proved elusive, however, with researchers so far able to conclude only that leptin plays some role in obesity in some people in some cases. Benner says examining the evolutionary history of the leptin gene family reveals that it evolved quickly between 40 million and 50 million years ago - after the divergence of rodents and primates in the lineage leading to human-like apes. That suggests its function in the body changed quickly as well, just as apes were moving to the top of the food chain. "Different biochemistry in humans and mice reflects their different strategies for survival," Benner says. "Correlating events in the genetic record with events in the fossil record suggested that leptin in humans may not play the same role as leptin in mice." This approach may prove useful for many other diseases, including Alzheimer's and osteoporosis, Benner says. The paper also for the first time correlates the contemporaneous evolution of different proteins, suggesting this approach could be used to find functional connections between different genes and proteins. These connections are important from the biomedical standpoint because they can open another door for scientists to control a disease, Benner says. For example, scientists may identify a protein key to Alzheimer's disease, but they may not be able to inactivate it because it also is key to essential life processes. Finding how it is triggered could reveal a way to turn off the part of the protein tied to Alzheimer's but leave the rest functional. Benner is the main author of the paper, with collaboration by M. Daniel Caraco, J. Michael Thomson and Eric Gaucher. Steven Benner, benner@chem.ufl.edu by Aaron Hoover Micro Surveillance Planes Get Closer To Reality
A poster in Peter Ifju's lab depicts the Predator unmanned aircraft, which snaps spy photos and fires missiles at enemy combatants in Afghanistan. But at worktables cluttered with wiring, tools and glue, Ifju and his colleagues and students at the University of Florida are at work on a completely different kind of drone.Each Predator weighs 2,300 pounds, has a nearly 50-foot wingspan and costs $3.5 million. Ifju's planes, by contrast, can be measured in inches and cost in the hundreds of dollars. Whereas the Predator travels thousands of miles and remains aloft for hours, the micro-aerial vehicle is designed for brief, short-range flights controlled by infantry soldiers. The idea is that soldiers will carry several of the planes in their backpacks, deploying them as needed to scout for enemy troops that may be just out of sight. "Soldiers can't carry around large surveillance aircraft. But they do need to look over the next hill," says Ifju. Micro-aerial vehicles, or MAVs, have been a hot area of research for several years. But Ifju's work is distinguished by rapid progress. When he and the other UF researchers launched their efforts in 1997, the first plane they produced spanned 18 inches, weighed 10 ounces and was powered by a gasoline engine. Ifju's latest model is a 5-inch, battery-powered craft. Even fully loaded with battery, video camera and transmitter, the oval-shaped plane weighs less than 2 ounces - yet remains aloft for 10 minutes and has a range of half a mile. "Our goal has always been to build mission-capable airplanes," Ifju said. It is not far off. Ifju's planes have taken first place at the annual International Micro Air Vehicle Competition for the past four years, with his 5.5-inch video-equipped MAV setting a new record in April for surveying a target 600 meters, or nearly 2,000 feet, from the launch site. In other words, his planes are rapidly moving from science fiction to science fact. Military research agencies such as the Defense Advanced Research Project Agency have poured millions into micro-aerial vehicle research, much of it for unproven and exotic "flapping wing" technology. Ifju's planes, by contrast, are propeller-driven and funded in part by U.S. Army Special Operations, which hopes Ifju will develop a system that its soldiers can begin using within the next few years. The National Science Foundation, the U.S. Air Force and NASA's Langley Research Center also have contributed to the UF effort, providing a total of about $500,000 so far.
Flying the planes requires a remote control, antenna and other hardware. Ifju and his colleagues have made the equipment portable, fitting it into a medium-sized plastic suitcase along with a catapult-like launcher, portable video player/recorder and virtual reality "video glasses." Made by Sony, the glasses reproduce the feel of watching a 52-inch TV from about six feet away. When connected remotely to the plane's camera, the glasses in effect place the pilot directly onboard the plane. Ifju's biggest hurdle may be the laws of aerodynamics. Small airplanes are inherently less stable than large ones because they have much more drag than lift. As a result, flying the MAVs requires considerable skill. Yet for the planes to be really useful, any soldier should be able to fly them. To attack this problem, Ifju and colleague Michael Nechyba, an assistant professor of electrical engineering, are working on a computer program that will act as a kind of MAV autopilot. The program stabilizes the plane by identifying the horizon based on the information transmitted in the digital image. Preliminary tests have been promising. Peter Ifju, pgi@aero.ufl.edu by Aaron Hoover Blood Stem Cells Can Make New Blood Vessels In MiceStem cells found in the bone marrow of adult mice don't just evolve into key components of blood - they are able to build blood vessels. The discovery, announced in May by University of Florida scientists writing in the online edition of Nature Medicine, marks an important step in the quest to master diabetes, cancer, heart disease, sight-robbing retinal disorders and a multitude of other medical conditions. The bone marrow is a rich source of stem cells, long heralded for their ability to mature into red and white blood cells and platelets. And as the key ingredient in a bone marrow transplant, stem cells already play a critical role in treating blood-borne cancers like leukemia. The latest findings open new lines of inquiry for scientists scrutinizing diseases involving excessive blood vessel growth, diseased arteries or other circulatory troubles. Doctors might someday be able to use the knowledge to encourage growth of new blood vessels that could repair injured organs or tissues or, alternatively, stop such growth when it could harm rather than help, said Edward Scott, an associate professor of molecular genetics at the UF Shands Cancer Center and director of the Program in Stem Cell Biology at UF's College of Medicine. "We've shown for the first time that adult blood stem cells can function to make blood vessels. We now know blood vessels can be formed from a remote location - they're not just repaired at the site of injury," Scott said. "That means we can now explore how to block that from happening to try to prevent things like diabetic retinopathy or to enhance that activity to promote new blood vessel formation or blood vessel repair." UF scientists said the breakthrough came through studies of more than 100 mice that were models for the human eye condition known as retinopathy. This diabetes-related complication, the leading cause of adult blindness in the United States, is characterized by a degenerative process that injures existing blood vessels and triggers new capillaries to form as the body struggles to repair the light-sensing retina. These new capillaries actually foster additional damage. The mice received bone marrow transplants of stem cells harvested from adult mice that were genetically engineered to glow green. Researchers were able to trace the origin of new blood vessels that formed in the animals' retinas back to the stem cells they had infused because the vessels glowed green. "We were able to show that, essentially, the majority of the vessels being repaired in the eye were green," Scott said. "We could see entire green blood vessels, whole new capillary beds that were green." The researchers also transplanted some mice with a single green stem cell. "Everything that glowed green in those animals, therefore, came from a single cell," Scott said. "That was our formal proof that it was indeed a blood stem cell that was making these blood vessels." Scott's UF collaborators included Maria Grant, W. Stratford May and Ammon Peck. The study was funded by the National Institutes of Health, the Juvenile Diabetes Research Foundation and the Leukemia and Lymphoma Society of America. The process by which blood stem cells form new vessels is likely to be very complex. Future research will focus on defining every step. Researchers also will seek ways to harness the body's ability to repair itself. Edward Scott, escott@ufl.edu by Melanie Fridl Ross University Surpasses $437 Million In Research Awards
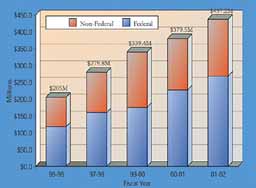
UF's Health Science Center accounted for just over half of the university's total, with its six colleges receiving a record $225.3 million, up 14 percent. Funding from the National Institutes of Health - Florida's largest source of research funding - rose 11.2 percent to $103.9 million. The university experienced a dramatic increase in funding from the National Science Foundation, climbing from $28.2 million in 2000-2001 to $39.2 million last year, a 39-percent jump. NSF funding supports research in the basic sciences, like physics and chemistry, engineering, and social sciences - like sociology and psychology. The College of Engineering saw a 34-percent increase to $67.7 million, thanks in part to a $15 million grant from NASA to lead a consortium of universities designing the next generation of reusable launch vehicle to replace the space shuttle. "We're very pleased NASA has identified the University of Florida as one of its centers of excellence," said Win Phillips, vice president for research and dean of the Graduate School. "This is a great accolade to our faculty who have been performing excellent research for NASA and for the country for years." Faculty in the College of Liberal Arts and Sciences brought in $38.1 million in 2001-02, up 13 percent over the previous year, while the Institute of Food and Agricultural Sciences brought in $69.5 million. Federal money accounts for 60 percent of UF's total research funding and this year reached $268.1 million, up more than 18 percent over last year. The university also continues to build its relationships with private funding agencies. Florida received nearly $54 million from industry sponsors in 2001-02 and another $48.3 million from foundations. "While publicly funded research, particularly from the federal government, is vital to our research enterprise, we have tried to achieve a more balanced ratio of public and private funding," Phillips said. Rat Liver Stem Cells Evolve Into Insulin-Producing CellsIn the latest breakthrough in the burgeoning field of stem cell biology, University of Florida scientists reported in June that adult rat liver stem cells can evolve into insulin-producing pancreatic cells, a finding that has implications for the future of diabetes research. Preliminary studies also show that the cells, when injected into diabetic mice, are able to reverse the animals' high blood-sugar levels, the researchers wrote in the June 4 online edition of Proceedings of the National Academy of Sciences. "Our major observation from this work is that adult stem cells from a non-pancreatic source can be pushed into becoming mature cells capable of producing and secreting insulin in response to glucose without genetically altering the mature cells," said UF pathologist Dr. Lijun Yang, who spearheaded the work, funded by the National Institute of Diabetes and Digestive and Kidney Diseases, in collaboration with UF scientists Ammon B. Peck and Bryon E. Petersen. The discoveries raise the possibility the liver could someday serve up a steady supply of cells with the potential to secrete insulin for patients with diabetes. "I think this study shows tremendous promise for the use of liver-derived cells to treat diabetes," said Dr. Diane Krause, an associate professor of laboratory medicine at Yale University School of Medicine. "It is important to note that this work represents a breakthrough that was built from extensive prior research on stem cells and the study of embryonic development." "In general, it has long been thought that blood stem cells could make only blood, and liver stem cells were born in the liver and could make only liver, and so forth," said Petersen, who is part of a multidisciplinary stem cell team that includes members of Florida's McKnight Brain Institute and Shands Cancer Center. "We were asking the question: Could liver stem cells be changed into a different cell type? Our latest findings basically show the flexibility of adult stem cells. We are now starting to understand how these cells work at the molecular and cellular levels and how to use this information to our advantage." Florida scientists isolated cells from the adult rat liver, then placed them in a high-sugar solution. The cells began to produce insulin, which liver cells don't normally do. Researchers subsequently implanted the cells into a small number of diabetic mice. Blood-sugar levels dropped to normal within 10 days in one mouse that had been given a high number of insulin-producing cells. Another two mice received much smaller numbers of cells and remained diabetic. Florida researchers currently are evaluating how long these cells will survive and function as insulin-producing cells, and precisely how effective they are at reversing diabetes in animal models, Yang said. "A major question is whether one injection of cells will last a lifetime or whether several injections will be needed," she said. "Another question is whether the change in the cell type is permanent or whether the cells can revert to liver cells. There are still many questions to be answered." by Melanie Fridl Ross Researchers Form Motor From Single DNA StrandThey are still many years away, but infinitesimal molecular motors that could radically improve manufacturing and medicine just took a step closer to reality. A University of Florida chemistry professor has made a "nanomotor" from a single DNA molecule. The motor, so small that hundreds of thousands could fit on the head of a pin, curls up and extends like an inchworm, said Weihong Tan, the principal investigator and lead author of an article about the motor in the April edition of the journal Nano Letters. While it is not the first such DNA motor, Tan said his nanomotor is the first to be built from a single molecule rather than several different DNA molecules. This makes it easier to use and edges such motors closer to real-life applications in the rapidly emerging field of bionanotechnology, Tan said. It is anticipated that nanomotors will play an active role in clinical treatment. For example, these ultra-small devices could be injected along with drugs that kill cancer cells or tumors, Tan said. When the drugs reach the disease site, the nanomotors would make the drug molecules attach and stick to the cancer cell membrane, Tan said. Perhaps more importantly, the motors' precision would give them the ability to prevent the drugs from attaching to noncancerous molecules or healthy parts of the body - eliminating the debilitating effects, for example, of chemotherapy drugs. Some scientists believe that nanomotors could also be used in so-called "test-tube manufacturing." This approach turns traditional manufacturing on its head. Where traditional manufacturing creates structures from existing materials or parts, test-tube manufacturing involves building structures from the smallest molecular or atomic components. Tan said the problem with the multiple-DNA strand motors already completed is they are very difficult to control because the pieces are so tiny - just tens of a billionth of a meter. Each DNA strand requires an energy source, which also reduces the motors' efficiency. With just one DNA strand, the UF nanomotor is easier to control and more efficient, Tan said. "Compared to other DNA motors, our nanomotor is more practical," Tan said. Tan and colleague Jianwei Li, a UF chemistry research associate, confirmed that the single-molecule motor worked by attaching a light-emitting organic molecule to one end and a light-quenching molecule to the other. When the motor extended, separating the quencher and emitter, the light went on. When it curled up, the light went out. Tan said it is difficult to predict when nanomotors, whether built from single or multiple molecules, will reach the stage when they can be used along with a drug or clinical treatment. He said the next step in his research is to coax his nanomotor to move a tiny particle from one place to another, demonstrating that it can perform a potentially useful task. Weihong Tan, tan@chem.ufl.edu by Aaron Hoover Australian Insect May Kill Invasive Melaleuca Trees
To slow - and hopefully stop - the spread of harmful Australian melaleuca trees in South Florida, researchers from the University of Florida and U.S. Department of Agriculture have released a tiny insect predator on the eastern edge of Everglades National Park. "After 16 years of research on natural or biological controls for the invasive tree species, we are releasing the melaleuca psyllid to help control or eradicate melaleuca," said Susan Wineriter, a senior agricultural biologist with UF's Institute of Food and Agricultural Sciences in Gainesville. "The psyllid, about the size of a gnat or small ant, feeds on melaleuca's clear sap, severely damaging seedlings," she said. "It's not a threat to people, animals or other plants." The melaleuca psyllid (Boreioglycaspis melaleucae) is the second beneficial insect imported from Australia to control melaleuca. In 1997, a weevil (Oxyops vitiosa) was released and is helping control melaleuca by feeding on leaves and flower buds. Seed production has been reduced by about 50 percent on trees it attacks. "Unlike the weevil, which is restricted to dry habitats, the melaleuca psyllid can invade in any melaleuca habitat," Wineriter said. "We expect this will provide more effective control of the tree." The Australian tree species was introduced to Florida in the mid-1880s. Since then, it has spread quickly throughout South Florida, displacing native plant and animal communities, drying up wetlands, creating fire hazards and threatening the stability of the Everglades ecosystem, she said. Susan Wineriter, spitz@ufl.edu by Chuck Woods $2.2 Million Grant To Study Tortoise Respiratory DiseaseBuilding on 10 years of research into an upper respiratory tract disease that has devastated endangered tortoises across the United States, University of Florida scientists hope a new $2.2 million federal grant will help them better grasp how various chronic diseases spread in the animals, as well as in people.
"The tortoise is unique, as it has about the same life span as a human and reaches reproductive age at about the same time," said Mary Brown, a professor of pathobiology at UF's College of Veterinary Medicine and a principal investigator. "Lots of changes have occurred in the tortoise's habitat, many of which are human-induced. We are interested in learning more about how natural factors combine with human-induced ones, such as relocation and fire exclusion, and about how those relationships interact with biological and microbial factors to determine the incidence and spread of disease." In the first year of the new project, a team led by Brown and colleague Paul Klein will survey more than 700 tortoises at 30 Florida sites to determine population characteristics, habitat quality and upper respiratory tract disease status. The sites include state parks, water management areas, military reserves, state mitigation parks and private property. In subsequent years, they will focus on 12 of these sites using ecological, molecular and other diagnostic tools to determine the influence of human-induced factors in disease spread and virulence. Researchers expect these multidisciplinary approaches to shed light on how respiratory disease in its various stages affects gopher tortoise populations. They also hope to develop mathematical models to predict the effect these multiple, complex factors have on disease spread. "Infectious diseases are an ever-present risk to wildlife, particularly during situations in which animals are removed from their natural habitats for captive breeding programs or during conditions of stress," Brown said. "This is even more important when the species concerned is a keystone species, such as the Florida gopher tortoise, that is critical to ecosystem health." As many as 360 animal species depend on the gopher tortoise for survival, including other threatened species such as the indigo snake. "Without the gopher tortoise, the biological diversity of upland habitats would be greatly diminished," said Klein, a professor and comparative immunologist in the UF College of Medicine who has a joint appointment in the veterinary college's Department of Pathobiology. "Furthermore, in a long-lived species that does not attain reproductive maturity for 10 to 20 years, a single catastrophic event such as a disease epidemic could reduce a population to the point that recovery would be extremely difficult." Such an event has happened to the threatened desert tortoise of the American Southwest, according to Kristin Berry, a biologist with the U.S. Geological Survey who first brought the respiratory disease, called mycoplasmosis, to the attention of the UF group a decade ago. "We have experienced catastrophic declines," Berry said. "We have lost at least 90 percent of our breeding tortoises in some populations, and in some of these groups, mycoplasmosis has played a role. It's going to take decades, if not centuries, for us to see recovery." Brown, Klein and others at UF who have studied this disease in both the gopher and desert tortoises, were first to identify the mycoplasma bacteria as the disease-causing agent. With a $150,000 grant from the Walt Disney Company, the group amassed data on several key populations of the gopher tortoise in Florida. "That work was an important foundation for the current research," Brown said. "It is exciting that the National Science Foundation is realizing the role of disease in the ecology of wildlife," Brown said. "They are recognizing that, in general, microbial infections in wildlife populations could have an impact on human populations, and that understanding how the disease spreads and what factors affect microbial virulence is very important." Mary B. Brown, mbbrown@nervm.nerdc.ufl.edu by Sarah Carey Fossil Suggests Early Flowers Grew UnderwaterThe world's oldest known flower never bloomed, but it has opened scientific questioning into whether all of today's flowering plants had their origins from beneath ancient waters, says a University of Florida researcher.
Although it had no petals, there is no question it was a flowering plant because of the presence of seeds enclosed in an immature fruit, a trait separating flowering plants from all other seed plants, he said. The discovery is important because it provides clues about how these now-extinct ancestors evolved into modern living flowering plants, said Dilcher, whose research is supported by the National Science Foundation. "Flowering plants are the dominant vegetation in the world today," he said. "They're the basic food crop and fiber source for the world's population. It's useful for us to understand the relationships among flowering plants, especially in this day of molecular genetic manipulations." The plant was about 20 inches high with thin stems stretching up in the water to the surface, with its pollen and seed organs extending above the water, Dilcher said. The seeds probably dispersed in the water, floated up along the shore and germinated in shallow water, he added. "The mysteries of the origin and radiation of the flowering plants remain among the greatest dilemmas facing paleontology and evolutionary biology," said William L. Crepet, professor and chair of the Department of Plant Biology at Cornell University. "This fossil represents the first evidence of an angiosperm that is basal to all other angiosperms, yet that does not fit within any modern taxonomic group of angiosperms - this makes it one of, if not the most important fossil flowering plants ever reported." The find was made in a fossil-rich section of the Yixian Formation northeast of Beijing, China, by local farmers who gave it to one of the paper's coauthors. It is much more complete than one found at a nearby site four years ago, which Dilcher also studied. "After having only a fragment and trying to imagine what the whole plant was like, it was a great surprise to find leaves typical of a plant that lived underwater, with characteristics very unique to flowering plants at such an early age in their history," he said. What distinguishes the fossil as an aquatic plant are its dissected leaves, Dilcher said. Some of the leaves at the base are quite branched, typical of underwater plants. Further proof of the flower's watery existence comes in the form of fish fossils found mixed in together with the fossil plants, he said. Dilcher, along with Ge Sun, a geologist at Jilin University in Changchun, China, and other researchers propose Archaefructaceae as the new basal angiosperm family for this ancient specimen. "These are the earliest, most complete remains of flowering plants yet discovered," Dilcher said. "What's spectacular about these fossils is that all parts of the plant are present, including the roots, leaves and reproductive organs. There's been some debate about whether the first flowering plants were woody or herbaceous. Now we see that they were herbaceous." David Dilcher, dilcher@flmnh.ufl.edu by Cathy Keen Formula Saves Airlines Money, Improves Service
A team composed of researchers from Florida, the Massachusetts Institute of Technology and United Airlines created a mathematical model that helps the airline identify the most profitable assignment of planes to flight legs and determine through connections between flights. The model, which relies on complex algorithms, is predicted to save the airline as much as $25 million annually and improve customer satisfaction once it is fully implemented. Each day, several hundred thousand passengers travel among hundreds of cities nationwide, with each city-to-city flight known as a "leg." Airline fleets consist of different types of planes with varying seating capacities. The airlines' goal is to assign the right plane to each flight leg while simultaneously determining the daily route of each plane - and minimizing its cost of operation. It's a gigantic scheduling problem. United Airlines' fleet, for instance, consists of more than 490 planes spanning nine different plane types from commuters to 747s. These planes usually fly an average of five flight legs per day or more than 1,700 daily flight legs. The problem is too complex to be solved manually, so airlines use mathematical models that computers solve. When assigning planes to flight legs, one model, known as the "fleet assignment model," matches the demand for each leg with the planes' seating capacities. When planes and flight legs have been matched, the airlines then take pairs of consecutive flight legs, such as Boston to Atlanta and Atlanta to Austin, and make them "through" flights using the "through assignment model." Passengers are willing to pay extra for these flights, because they don't have to change planes, so airlines want to make as many of them available as possible. United realized that by combining the two models it could improve profitability. But the combined model was too difficult to be solved with the available technology. In a research project sponsored through a $500,000 grant from the National Science Foundation, Ravindra Ahuja, a professor of industrial and systems engineering at Florida, and James Orlin, a business professor at MIT, developed a novel technique to solve such difficult problems quickly. Known as "Very Large-Scale Neighborhood Search," the technique starts with a feasible solution for a problem and repeatedly replaces it with an improved "neighbor" solution. The NSF grant enabled the Ahuja-Orlin team to develop a method that allowed them to test trillions of neighbors and determine the best solution in a fraction of a second on a typical personal computer. United then asked the team to develop an application of the technique to solve its combined scheduling model. The software used the solution produced by United's current software as the starting solution. It was improved by using the neighborhood search that takes less than five seconds to complete. The research was sponsored by United's Research and Development Department. Two doctoral students, Dushyant Sharma from MIT and Jian Liu from UF, also participated. "We are excited about this new technology," said Raj Sivakumar, director of research and development at United. "It will be a key component of our overall strategy for flight scheduling." Ravindra Ahuja, ahuja@ufl.edu by Aaron Hoover |
 Clues for using the sequence of the human genome to diagnose and treat diseases may lie in our distant past, says a University of Florida professor.
Clues for using the sequence of the human genome to diagnose and treat diseases may lie in our distant past, says a University of Florida professor.

 The skeleton of each plane consists of strands of carbon fiber. Latex rubber stretched over this skeleton forms the wings. A small lithium-polymer battery, similar to that used in cellular phones, and a video transmitter fit into the fuselage, which is made of clear plastic film. The pencil eraser-sized video camera lens pokes from the front of the fuselage. Each plane takes about five hours to build and costs $700, but $450 is for the video transmitter alone, Ifju said.
The skeleton of each plane consists of strands of carbon fiber. Latex rubber stretched over this skeleton forms the wings. A small lithium-polymer battery, similar to that used in cellular phones, and a video transmitter fit into the fuselage, which is made of clear plastic film. The pencil eraser-sized video camera lens pokes from the front of the fuselage. Each plane takes about five hours to build and costs $700, but $450 is for the video transmitter alone, Ifju said.

 Funded by the National Science Foundation, the Florida-based project is one of the largest of its kind ever awarded for research involving wild animal disease as a model for understanding the impact not only on humans but also on the entire ecosystem.
Funded by the National Science Foundation, the Florida-based project is one of the largest of its kind ever awarded for research involving wild animal disease as a model for understanding the impact not only on humans but also on the entire ecosystem.
 The newly discovered remains of the oldest, most complete flowering plant show it lived at least 125 million years ago and likely was an underwater plant, said David Dilcher, a graduate research professor of paleobotany at Florida. The discovery was reported in April in the journal Science.
The newly discovered remains of the oldest, most complete flowering plant show it lived at least 125 million years ago and likely was an underwater plant, said David Dilcher, a graduate research professor of paleobotany at Florida. The discovery was reported in April in the journal Science.
 A University of Florida-led research team has developed a new mathematical approach to airline scheduling that could lead to substantial savings for carriers.
A University of Florida-led research team has developed a new mathematical approach to airline scheduling that could lead to substantial savings for carriers.